Description

Source: MedicalExpress
Disclaimer: Copyright infringement not intended.
Context
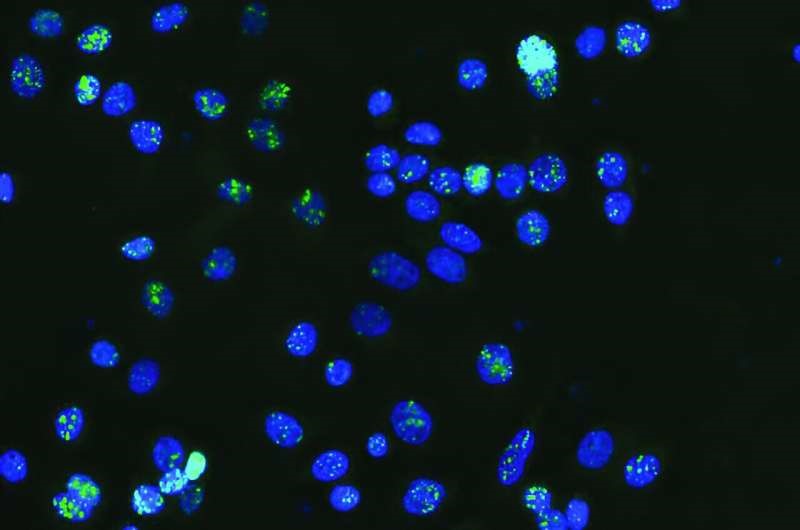
- Researchers at the Garvan Institute of Medical Research have identified potential cancer drivers in the non-coding regions of DNA, commonly referred to as 'junk' DNA.
- Utilizing artificial intelligence, the study opens new avenues for cancer diagnosis and treatment by exploring previously overlooked areas of the genome.
Details
Key Findings
- The study focuses on the 98% of the human genome that does not encode proteins. Historically considered non-functional, this 'junk' DNA may hold crucial insights into cancer development and progression.
- Discovering mutations in these regions could lead to early cancer diagnosis and the development of new treatments that are effective across multiple cancer types.
- The researchers concentrated on mutations affecting CTCF binding sites. CTCF is a protein that helps fold DNA into specific 3D shapes, crucial for gene regulation.
- Persistent CTCF binding sites act as 'anchors' in the genome, maintaining the 3D structure and regulating gene expression.
Junk DNA
- Junk DNArefers to regions of DNA that are non-coding, meaning they do not code for proteins. Initially, these regions were thought to have no functional purpose, hence the term "junk."
- However, recent research has revealed that many of these sequences play crucial roles in gene regulation, chromosomal structure, and genome evolution.
Types of Non-Coding DNA
- Introns: Non-coding sequences within genes that are removed during RNA splicing.
- Regulatory Sequences:
- Promoters: Regions where RNA polymerase binds to initiate transcription.
- Enhancers: Sequences that can increase the transcription of associated genes.
- Repetitive DNA:
- Satellite DNA: Consists of large blocks of tandemly repeated sequences.
- Minisatellites and Microsatellites: Shorter repetitive sequences, often used in genetic fingerprinting.
- Transposable Elements: DNA sequences that can change their position within the genome, potentially creating or reversing mutations.

Functions of Non-Coding DNA
- Gene Regulation: Many non-coding regions contain regulatory elements that control when and where genes are expressed. This includes enhancers, silencers, and insulators.
- Structural Roles: Non-coding DNA contributes to the physical architecture of the genome. For example, telomeres and centromeres consist of repetitive non-coding sequences essential for chromosome stability and segregation.
- Genome Evolution: Non-coding DNA can drive evolution by providing raw material for genetic variation. Transposable elements, for instance, can create new regulatory networks or gene fusions.
- RNA Molecules: Some non-coding regions are transcribed into functional RNA molecules (e.g., rRNA, tRNA, and various types of regulatory RNAs such as miRNA and lncRNA).
Research and Applications
- ENCODE Project: The Encyclopedia of DNA Elements (ENCODE) project aims to identify all functional elements in the human genome, revealing that a significant portion of non-coding DNA has biochemical functions.
- Disease Association: Mutations in non-coding regions can be associated with diseases. For example, variations in enhancers or silencers can lead to misregulation of gene expression, contributing to cancer, genetic disorders, and other conditions.
- Biotechnological Applications: Understanding non-coding DNA can improve gene therapy, personalized medicine, and the development of new therapeutic strategies targeting regulatory elements.
Sources:
MedicalExpress
|
PRACTICE QUESTION
Q: The term "junk DNA" is a misnomer in light of modern genomics. While much of the genome does not code for proteins, non-coding DNA plays essential roles in gene regulation, chromosomal architecture, and evolutionary processes. Explain. (250 Words)
|